Difference between revisions of "Knowledgebase mockups"
(→Search summary results) |
(→Comments) |
||
| Line 9: | Line 9: | ||
====Comments==== | ====Comments==== | ||
*From Paula: start off with taxon tree semi-collapsed (at genus level). | *From Paula: start off with taxon tree semi-collapsed (at genus level). | ||
| − | *Put the number of taxa for which there are annotations in parens behind, e.g., the genus name. | + | *From Paula: Put the number of taxa for which there are annotations in parens behind, e.g., the genus name. |
===Search summary results=== | ===Search summary results=== | ||
Revision as of 18:11, 15 May 2009
Contents
Grouping data by using an ontology tree view
Phenotype results
This would be an enhancement to the results currently displayed on a page like http://kb.phenoscape.org/phenotype/evo/TTO:10930/TAO:0000326/PATO:0000070/
The nearest common ancestor term of the annotated taxon terms is used as the root of the tree display. The only leaf terms displayed are those with annotations. Internal nodes show the union of the annotations to their descendant nodes.
Comments
- From Paula: start off with taxon tree semi-collapsed (at genus level).
- From Paula: Put the number of taxa for which there are annotations in parens behind, e.g., the genus name.
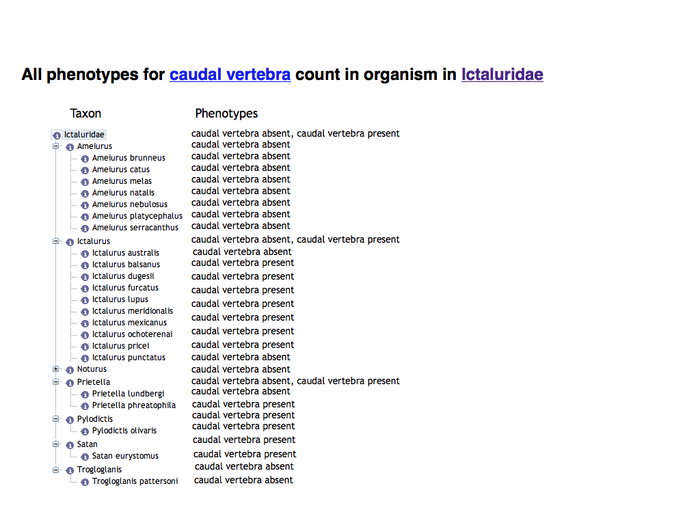
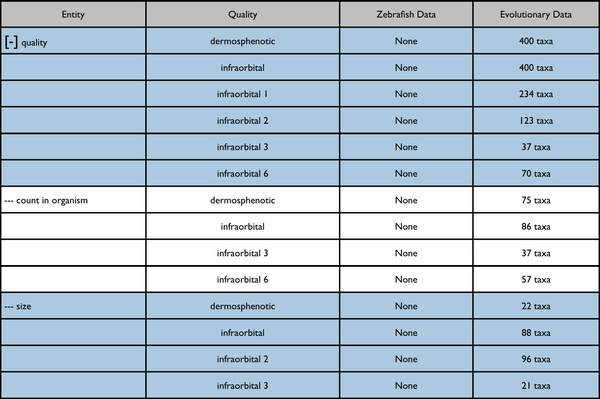
Search summary results
This would be an enhancement to the Phenotypes results currently displayed on a page like http://kb.phenoscape.org/search/anatomy/TAO:0000376
The leftmost column is intended to represent a disclosable tree display. The nearest common ancestor of the represented annotated anatomy terms is used as the root of the tree display. Each anatomy term has a flat list of the qualities annotated for it. Don't pay too much attention to the dummy numbers given in these results.
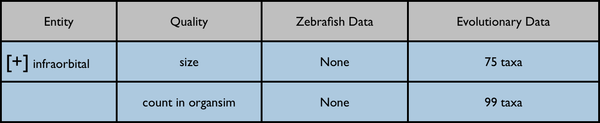
Root anatomy term "infraorbital" collapsed:
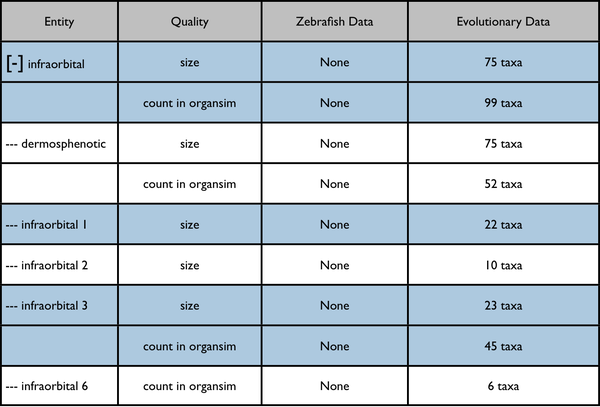
Root anatomy term "infraorbital" expanded:
Possible further enhancement to page capabilities - redisplay data on a quality tree. The included results would be somewhat different.
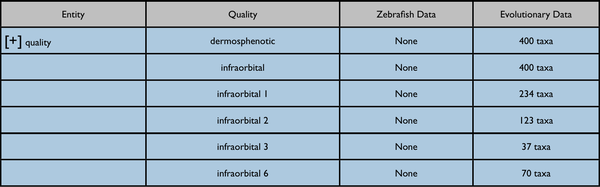
Nearest quality ancestor collapsed:
Quality ancestor expanded:
Comments
- From Paula: Rename 'Entity' 'Anatomy' as per other mockups for consistency http://kb.phenoscape.org/search/anatomy/TAO:0000376
- From Paula: swap column headings Entity-Quality on Quality tree table (below ' Nearest quality ancestor collapsed" text)
Display of citation and free-text basis for annotations
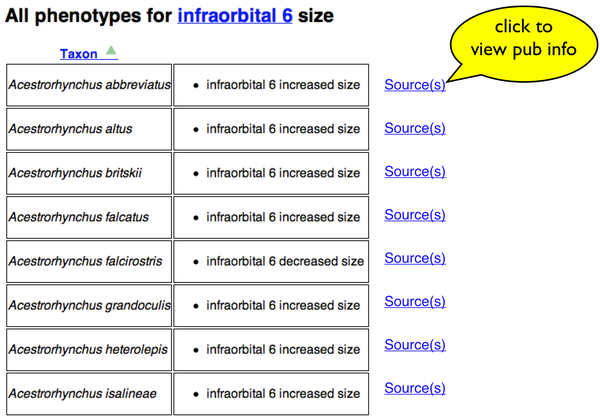
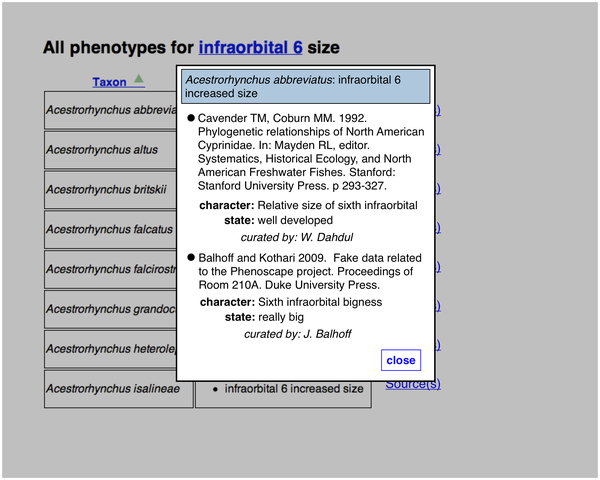
Here is a list of specific phenotype annotations resulting from a search. There is an added "Source(s)" link next to each one. The yellow bubble is not part of the page.
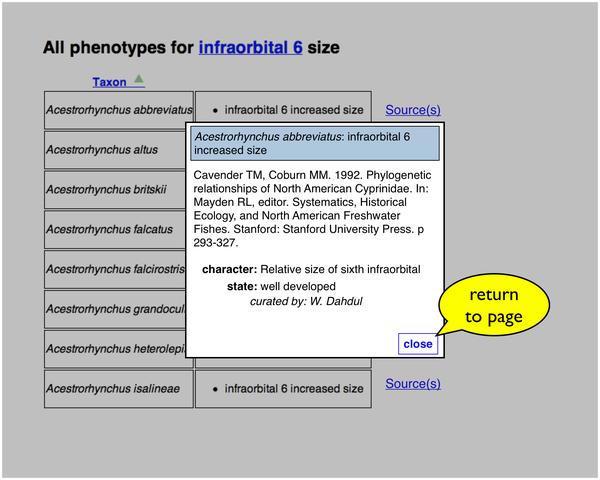
If the user clicks a "Source(s)" link, the citation information for a particular taxon-phenotype annotation is retrieved and displayed in a page overlay.
A particular taxon-phenotype annotation may have been asserted from more than one publication:
Simple display of homology information
Anatomy pages can make use of homology information from the database to list possible homologous terms. This mockup adds a list of homologous terms to the term info panel. The user can click one of these term links to go to the anatomy page about that term.