Project Plan
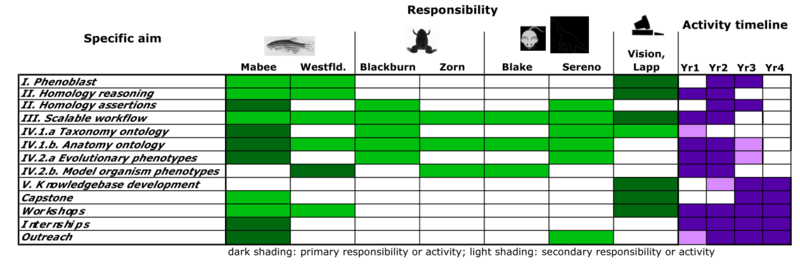
Much of the schedule below is based on the 1 July 2011 funded NSF grant. Our development schedule is revised as shown in the figure below in concert with budget cuts. Relative to the original proposal the support for curation, particularly in the model organism databases, is significantly reduced. This will impact our ability to respond as any problems arise with integration of the evolutionary data and somewhat compromise our ability to rigorously evaluate the tools being developed. Additionally, we will scale back the development goals for the semantic similarity search engine, focussing our efforts on achieving scalability and speedup. We have also frontloaded the plans for execution of the NLP work to achieve a scalable workflow as early as possible in the process.
Goals in progress, coming up, and scheduled
<embedurl>https://trello.com/b/xYkl0qmC</embedurl>
Capstone
As a capstone, in years 3 and 4 of the project, we will validate the capabilities of the above suite of tools by testing how well known developmental pathways for the well-studied fin/limb skeletal transition in vertebrate evolution are identified and how well it scales to a datastore containing billions of phenotypes.
NSF Phenoscape project abstracts
DBI 1062404 and 1062542: Collaborative research: ABI Development: Ontology-enabled reasoning across phenotypes from evolution and model organisms
1. Technical description of the project.
An award is made to the University of South Dakota and the University of North Carolina to develop ontology-driven tools for machine reasoning over large volumes of phenotype data. A fast semantic similarity engine will be developed to allow searches for evolutionary transitions and mutant genes characterized by similar phenotypic profiles. An ontological framework for reasoning over homology will be developed to allow rigorous reasoning over evolutionary diverse lineages. Natural language processing tools will be developed to improve upon the efficiency of mining phenotype data from the literature and improving data consistency. This suite of tools will be tested on a large number of skeletal phenotypes from diverse fossil and modern vertebrates. Taxonomic and anatomical ontologies for vertebrates will be augmented and hypotheses of anatomical homology formally encoded. The ontologies and software tools, together with phenotypes extracted from the vertebrate systematic literature, will be integrated in the knowledgebase with genetic and phenotype data from three vertebrate model organisms: zebrafish (Danio rerio), African clawed frog (Xenopus laevis), and mouse (Mus musculus). The knowledgebase will be exposed to generic reasoners using semantic web standards. The system will be validated by its success in retrieving candidate genes for the well-studied vertebrate fin-limb transition and other major events in skeletal evolution.
2. Non-technical explanation of the project's broader significance and importance.
Human-readable descriptions of “phenotypic” properties such as anatomy and behavior are not well-suited to computational analysis. Yet, in evolutionary biology, genetics and development, computational assistance is necessary to discover patterns within the enormous volumes of descriptive phenotype data that are being reported in the literature and in online databases. Ontologies are structured, controlled vocabularies that can be applied to collections of descriptive data to permit logical reasoning to be used. Using the evolutionary transition from fins to limbs as a test system, this project will develop ontologically-aware software that allows users to discover similar sets of phenotypes for different taxa or mutant genes within large and diverse datasets. The evolutionary breadth of the test data requires the development of a rigorous framework for reasoning over hypotheses of homology. Another goal is to develop and evaluate natural language processing tools for efficiently capturing ontological descriptions of phenotype from the descriptions available in the published literature. Phenotype data from the systematic literature for both extinct and extant vertebrates will be combined with mutant phenotype data from three vertebrate genetic models: zebrafish (Danio rerio), frog (Xenopus laevis), and mouse (Mus musculus). The suite of tools will be validated by recovering developmental genetic pathways that underlie the evolutionary transition from fin to limb in vertebrates, and refined by iterative testing with domain bioinformaticians on the project and biologists from the broader user community.
3. Indicate how your project addresses criteria specific to Development
A broad community of users will participate through the lifecycle of this project in the development of community standards and resources for the interoperability and computability of phenotypic knowledge. This will be achieved through workshops, usability testing sessions, and coordination with key research networks. Stakeholder ownership will be enhanced by rapid and open release of a variety of products that we anticipate to be of immediate and enduring value to the greater biology community, including tools for streamlining data curation and performing large-scale semantic similarity searches, high quality vertebrate taxonomy and anatomy ontologies, and standards for reasoning over homology. We will provide a unique training environment for students, postdocs and summer interns, including Native Americans through outreach at the University of South Dakota and minority and female students though a collaboration with Project Exploration at the University of Chicago. Project progress and outcomes will be disseminated through both traditional and online outlets for scholarly communication (including blog posts at mailing lists); the primary web presence will be at https://www.phenoscape.org/wiki/.