Phenote is being used for phenotype annotation of mutants within model-organism projects. It allows the creation of a simple list of phenotype statements, rather than a species-by-character matrix.
Here is sample data in the default configuration:
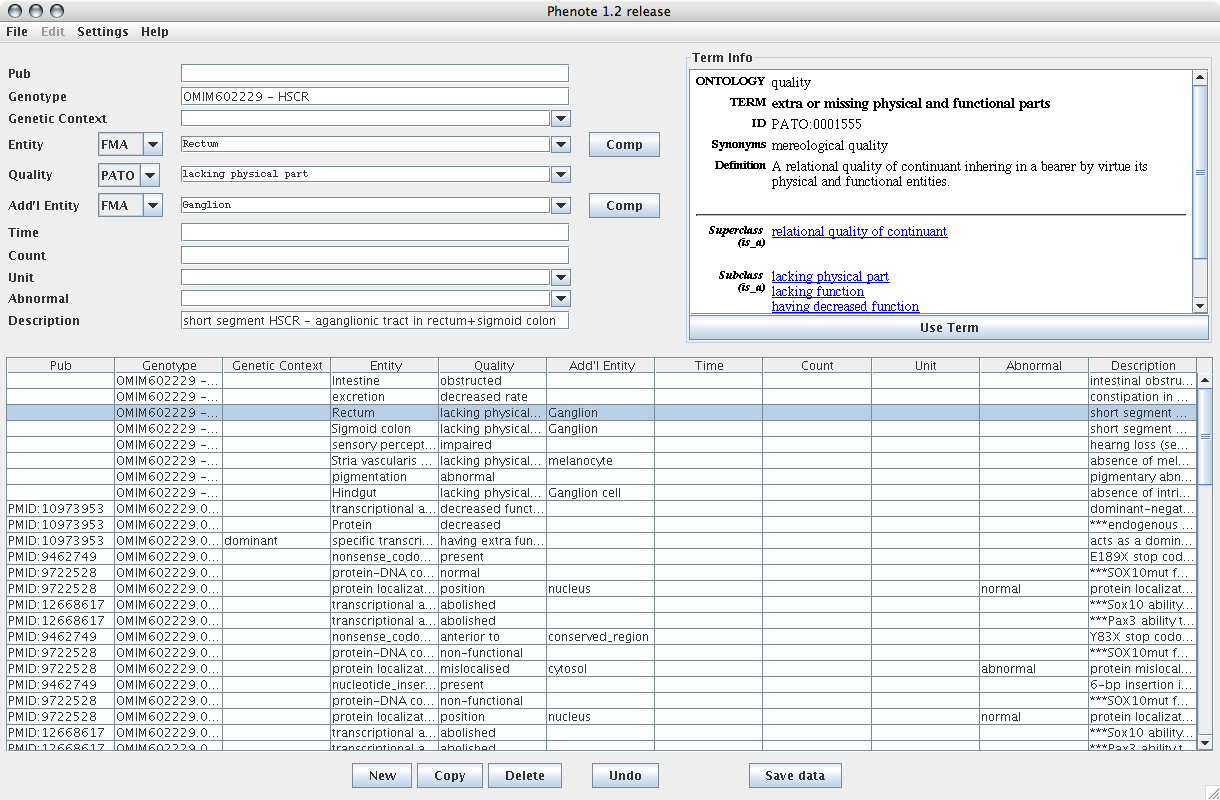
The input fields in Phenote are highly configurable. Here is a sample of how Phenote might be configured for the PhenoScape project:
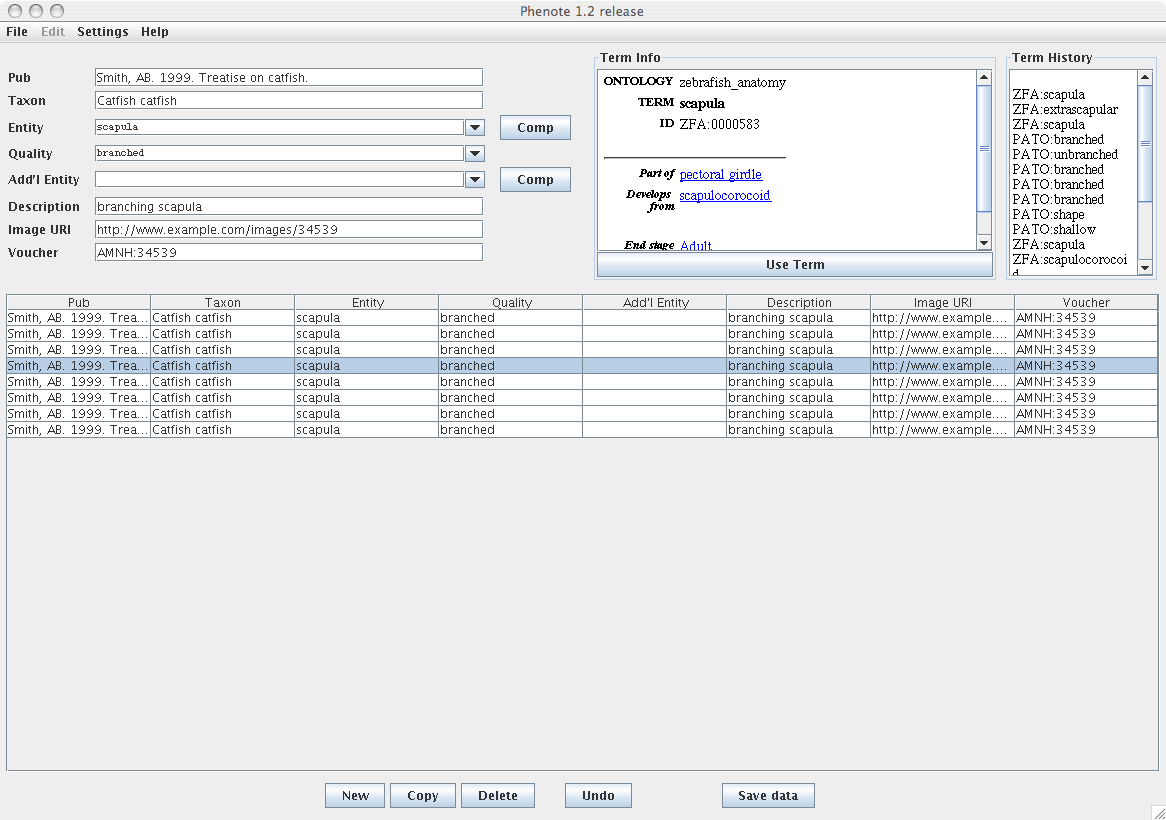
This configuration file can be downloaded here. Phenote configurations are described in the documentation.
This interface (“list of states”) may not be optimal for the PhenoScape project, especially if a set of species are expected to be coded for a given set of characters.