Difference between revisions of "Knowledgebase mockups (Nov 2009)"
From phenoscape
(→Publication results on anatomy term search pages, revision 1) |
(→Comments:) |
||
| Line 11: | Line 11: | ||
=====Comments:===== | =====Comments:===== | ||
* John likes separate column for publications (as in previous revision), maybe add 'KB Publications' vs. book icon in taxon column. | * John likes separate column for publications (as in previous revision), maybe add 'KB Publications' vs. book icon in taxon column. | ||
| + | * Tree page, source data box: will need link to Pub page 3, below. Case # | ||
====List of publications, revision 1==== | ====List of publications, revision 1==== | ||
Revision as of 15:21, 13 January 2010
Contents
Publications
Revision 1
Revision 1 posted on 12/16/09 by Wasila based on amphibanat feedback, Phenoscape conference call (11/20/09), discussion with Paula (12/14/09), and conference call with John Lundberg (12/18/09). The Powerpoint file containing the revisions is on the webdav.
Publication results on anatomy term search pages, revision 1
Comments:
- John likes separate column for publications (as in previous revision), maybe add 'KB Publications' vs. book icon in taxon column.
- Tree page, source data box: will need link to Pub page 3, below. Case #
List of publications, revision 1
Publication detail page, revision 1
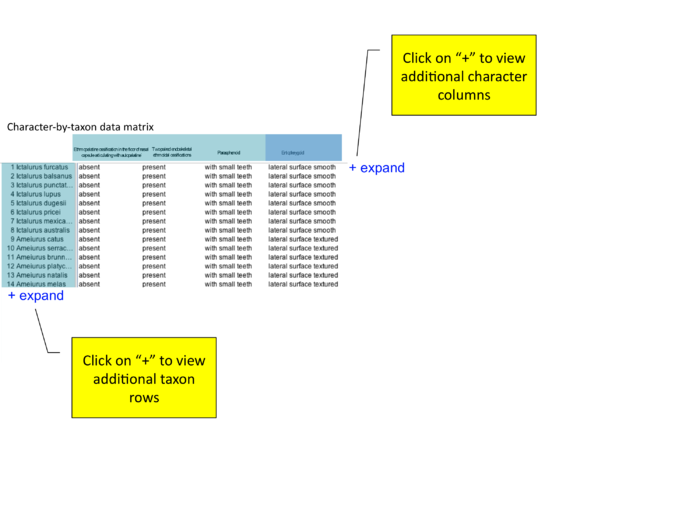
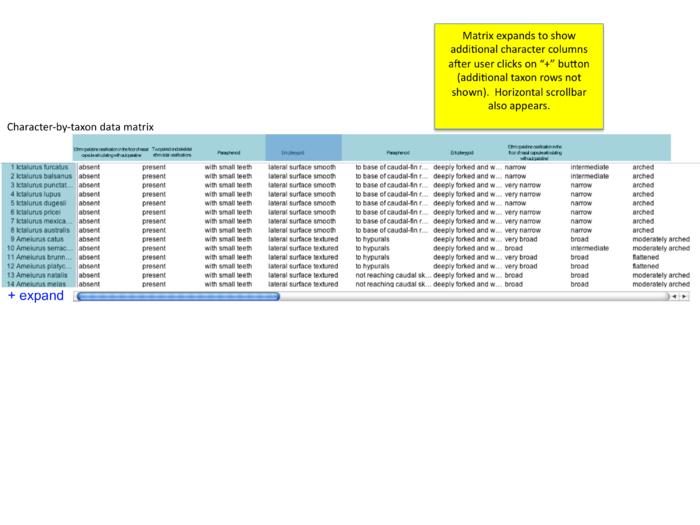
Additional character x taxon matrix mockups
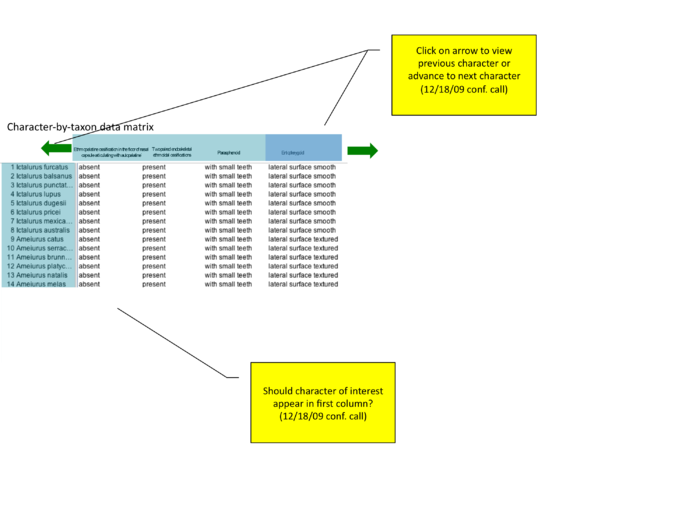
Based on conference call 12/18/09; mockups added on 12/21/09 by Wasila. "how to show character of interest? should entopterygoid be first column (even if author lists as #4 or #79)? or should there be a slider or 'next' arrow that allows users to view additional characters?"
collapse/expand all button
advance to next character arrow
Comments:
Data downloads: phenotypes for single entity across studies
From 12/18/09 conference call:
- John’s new use case: obtain data for a single entity across multiple studies. If want to do broad phylogenetic analysis of catfishes, need to look in detail in each pub of how entopterygoid is treated. Want to get just the entopterygoid by taxon data matrix.
- go back to anatomical term page. Need to get all data for entopterygoid (size, shape, etc.). Will the tree have 'entopterygoid' with 'ento, shape', 'ento, size' etc. beneath it? Will there be a sum/number of the total number of phenotypes for just entopterygoid? if, e.g. '800', click on it, and you would see a matrix with columns, size, shape, etc. with EQ in cells, but mouse-over to see free text. Can DOWNLOAD THIS SELECTION OF DATA ALONE in excel. In the excel file, you would have the free text, publication, and EQ? ...same thing as in genbank data with incomplete stuff, put in alignment program.
Original mockups
Integration of publication data across site.
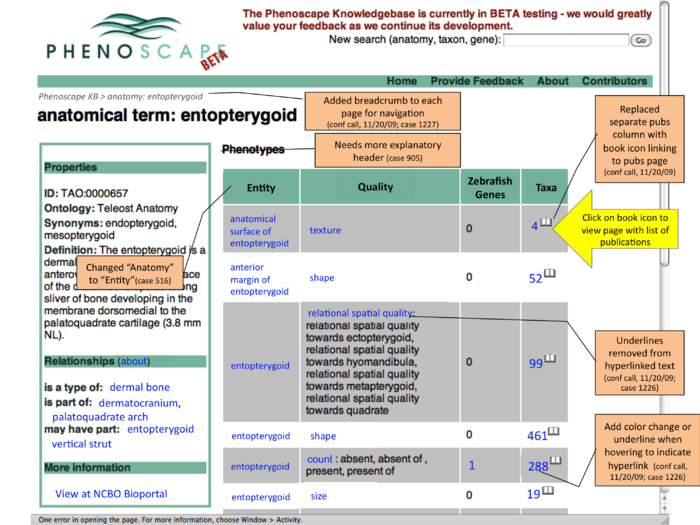
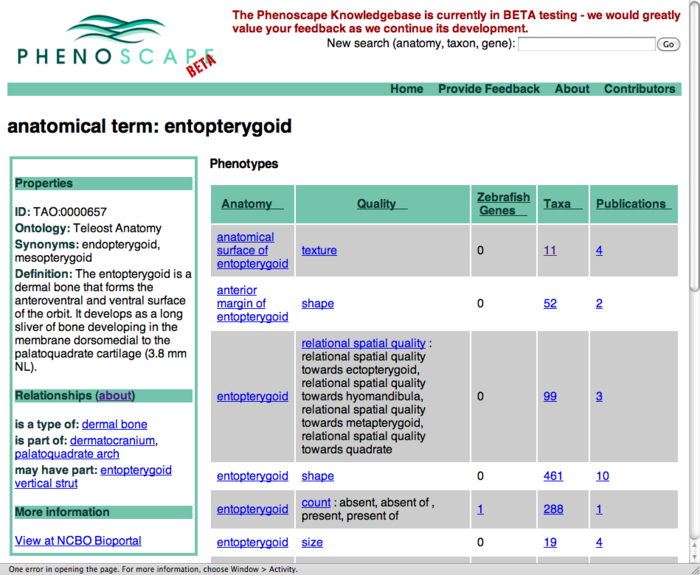
Publication results on term search pages
- this shows an entity page (entopterygoid) - could have similar layout on taxon and gene searches
- for each row (phenotype class) - see how many publication are linked to any EQs of that type
- click one of the count links to go to a page listing those publications
List of publications
- click one of the links to go to that publication's detail page
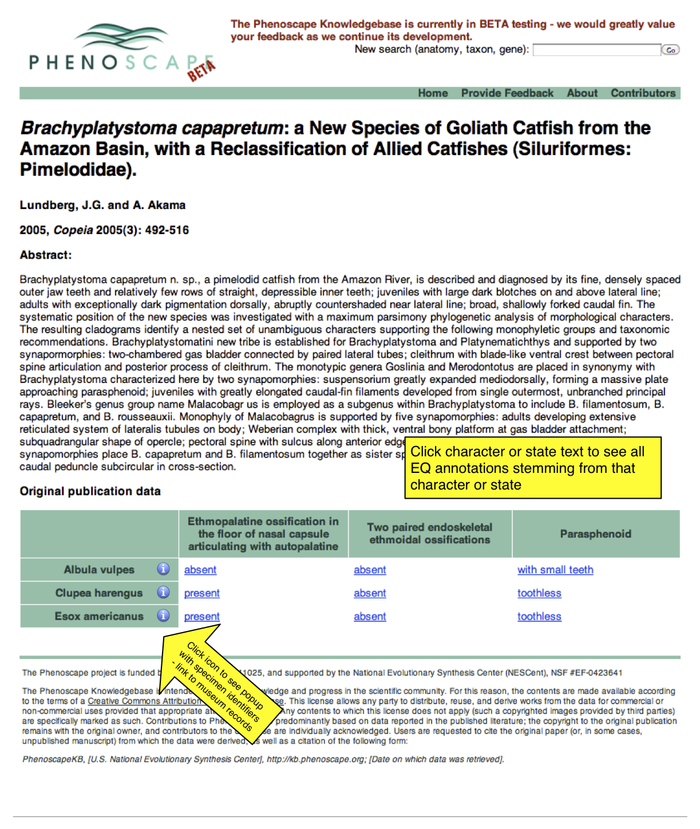
Publication detail page
- matrix will usually be much bigger - will be scrollable
- click column header or cell links to go to a phenotype details page showing all related EQs grouped by taxon (as on existing phenotype pages)
Comments
Tree mapping
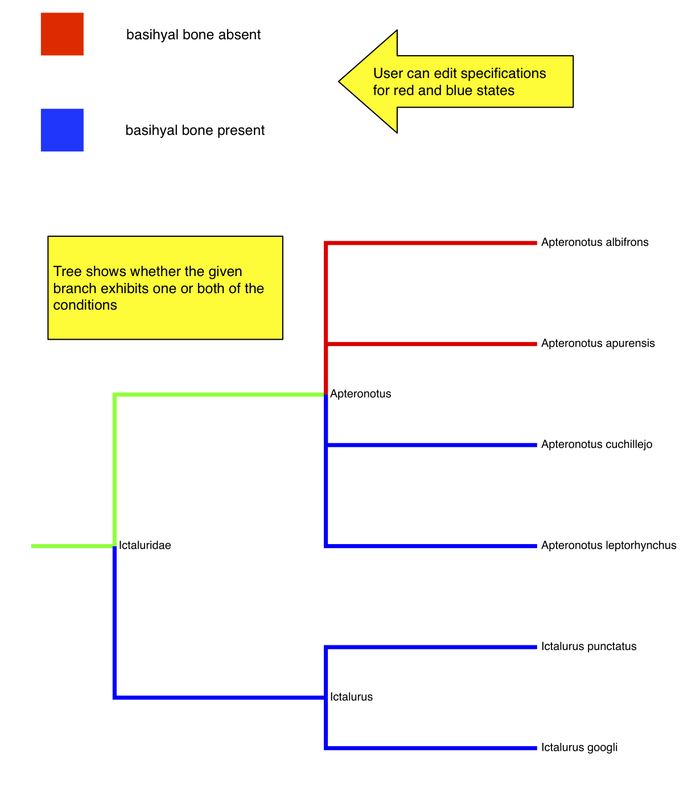
Map 2 phenotypes on tree
- user can enter phenotype specification for each "state"
- see on tree which branches exhibit one or both of the phenotypes
- Do users want the tree "pruned"? Or show branches without annotations also? could be a lot of taxa
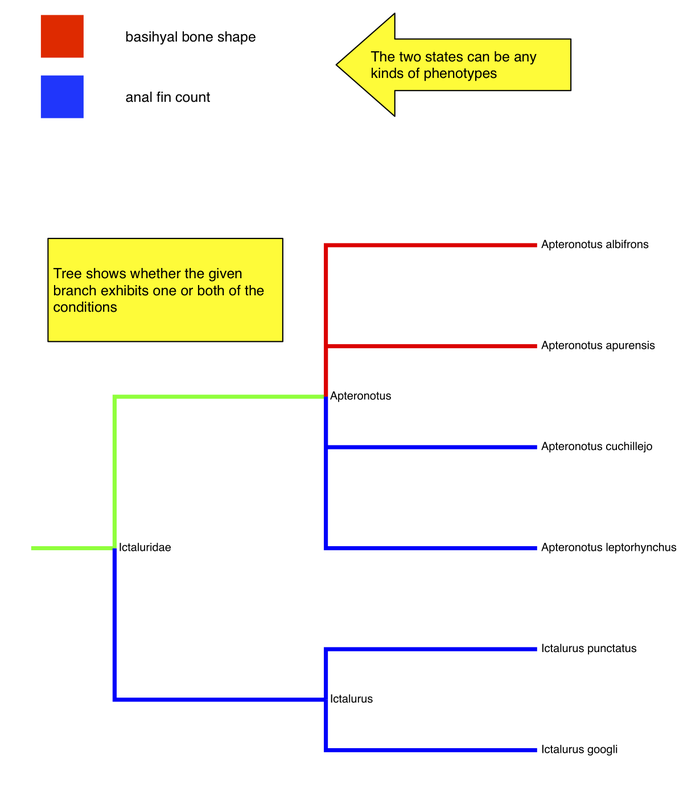
Same interface
- shows that the 2 phenotypes don't have to be obvious alternatives
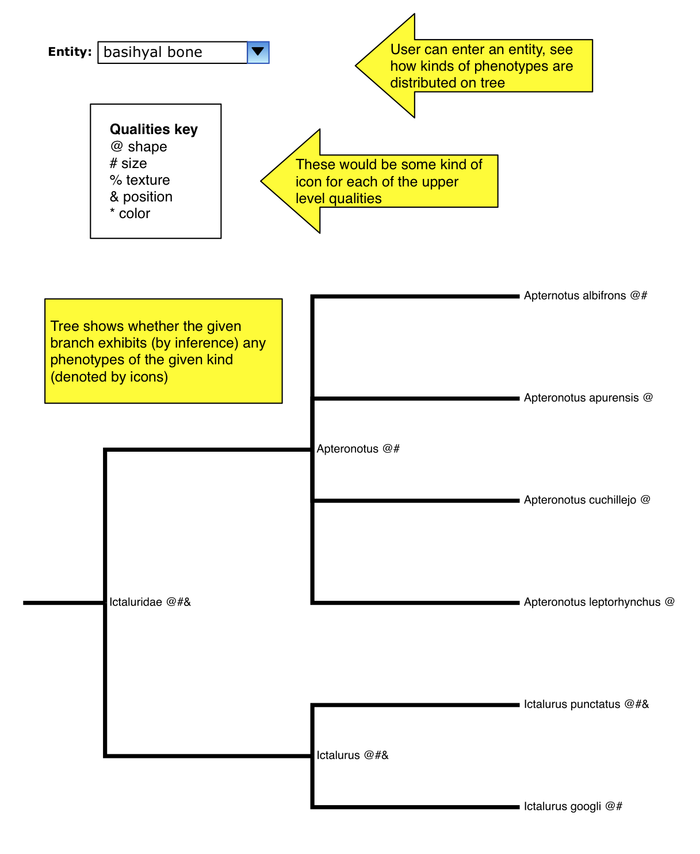
Map phenotypes of a given entity on tree
- user enters any entity term
- each of the standard upper-level PATO qualities has some icon
- each node of tree is annotated with icons for which kinds of phenotypes are exhibited by that node
- so if you just put in "bone", you could get a "shape" icon at a node for any bone shape phenotype (basihyal bone round, vertebra 1 flat, etc.)
Comments
Phenotype queries
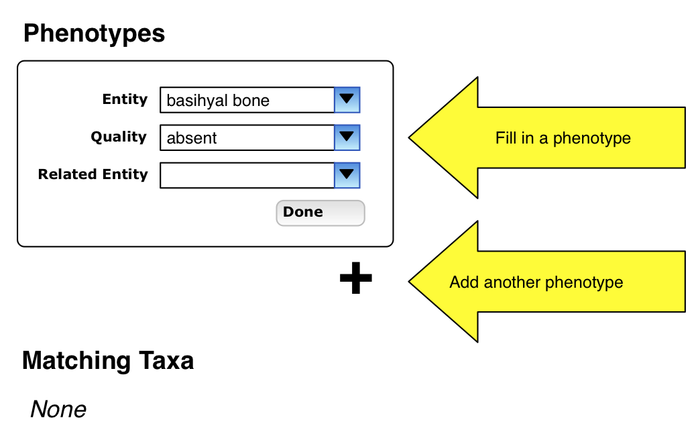
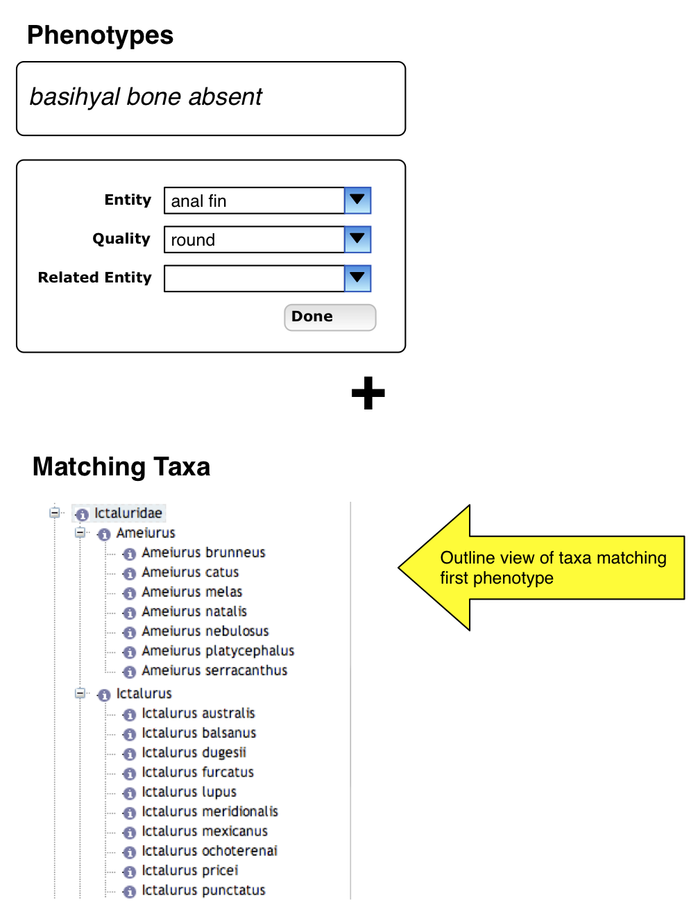
Entry of phenotype specification
- user enters a phenotype specification, presses Done
Additional phenotype specification
- bottom of page shows taxa which exhibit first phenotype
- user pressed "+" under first phenotype and can enter another
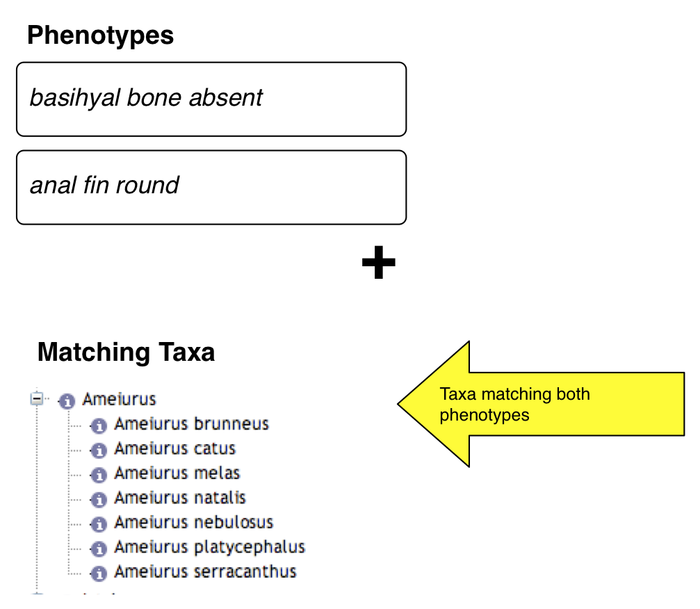
Results
- taxa exhibiting both phenotypes are shown
Comments
What should happen upon clicking taxon names? --jim 11:38, 18 November 2009 (EST)
- go to taxon's page?
- go to phenotype listing for that taxon and given phenotypes?