Difference between revisions of "Taxonomy ontology"
m (→Resources) |
|||
| Line 512: | Line 512: | ||
is_a: TTO:taxonomic_rank ! taxonomic_rank | is_a: TTO:taxonomic_rank ! taxonomic_rank | ||
</pre> | </pre> | ||
| + | |||
| + | |||
| + | Note: This page describes some design alternatives for taxonomic ontologies that refer to or include terms for ranks. If you are looking documentation for the Vocabulary of Ranks developed by Phenoscape, you will find it [[Taxonomic Rank Vocabulary | here]]. | ||
| + | |||
| + | A taxonomic rank ontology allows particular taxa to be treated as historical individuals instead of as universal types. | ||
| + | |||
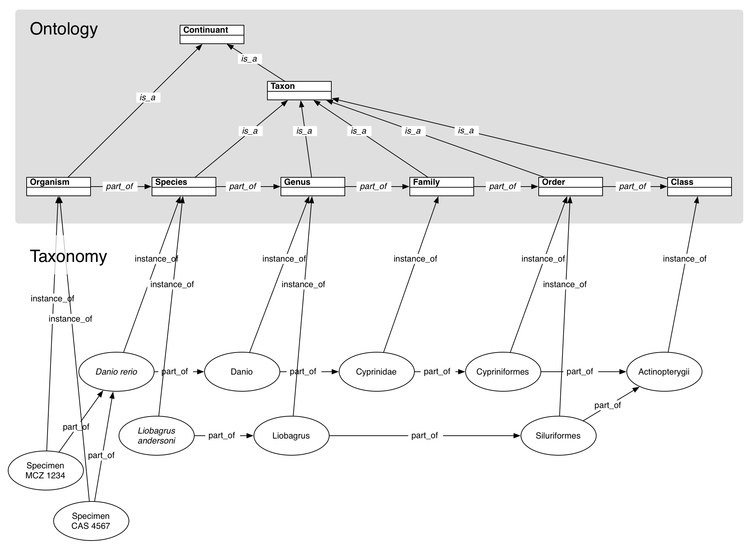
| + | [[Image:TaxonomicRankDiagram.png|750px|thumb|Taxonomic rank ontology and sample instance data. The ontology terms are surrounded by a grey background. The "Continuant" term would reside in an upper ontology such as [http://www.ifomis.org/bfo BFO]. The instance data constitute a particular taxonomy.]] | ||
| + | |||
| + | Previous attempts to represent taxonomy using ontology usually include taxonomic groups as classes in the ontology. Individual organisms are seen as instances of those universal types. There could be an ontology term Mammal, such that Primates and Rodents are more particular types of Mammals (''is_a'' descendants). Taxonomies are even often used as examples to help explain ontology inheritance to new users. This scheme fails to represent reality in several ways and, as this page will demonstrate, is even misleading. | ||
| + | |||
| + | ==The "traditional" view== | ||
| + | |||
| + | ===Inference with traditional taxonomy=== | ||
| + | One of the attractions of a traditional taxonomy for ontology builders that it can support the most common sort of reasoning for Description Logic ontologies. Although neither the TTO nor the NCBI taxonomy ontology attach character values to their classes, in principle such characters could be extracted from keys or from the taxonomic literature. With the addition of such properties, individuals can be classified by starting at the root of the taxonomy and traversing down the tree until either a terminal taxon (species) is reached or a node is reached where no descendant has the combination of characters displayed by the individual. For biologists, this operation is identical to use of an identification key. Such an operation is generally only defined for ''is_a'' links. | ||
| + | |||
| + | ===Problems with the "traditional" view=== | ||
| + | * The only criterion for inclusion within a taxonomic group is physical descent from a member of that group (we are assuming a taxonomy consistent with phylogeny). There are no universal properties one could ascribe to members of "Mammalia", besides things the members happen to share at this moment. "Hair" is often used an example of a Mammalian property; but of course its commonality is the result of its presence in the common ancestor of all mammals, and many mammals lack hair almost entirely. A mammal species that had no hair at all would nevertheless still be a mammal. Such reversals are common in phylogenies and can occur for both morphological properties (e.g., loss of legs in snakes) and molecular characters (e.g., inferred origin of pseudogenes). | ||
| + | * Descendant terms do not inherit (ontologically) all of the characteristics of their parents. For example, Rodentia has a different age and geographic range from Mammalia. | ||
| + | * Properties of a taxonomic group change through time: its geographic range, genetic and morphological diversity, etc. It can go extinct. These properties inhere in the group, not any one individual. | ||
| + | * Treating taxonomic groups as "classes" or "types" reinforces essentialist ideas of life forms, which evolutionary biology has completely overturned. Essentialist thought is a hindrance to understanding of the evolutionary process. | ||
| + | * Most contemporary treatments of the species concept (refs) treat them, and in many cases higher taxa, as individuals, possessing a history. | ||
| + | |||
| + | Instead of treating taxa as types in an ontology, we can treat individual taxa as instances of the type Taxon, or more specifically as instances of some taxonomic rank (see figure above). | ||
| + | |||
| + | ==Taxonomic Rank Ontology + Instance Graph of Taxonomy== | ||
| + | |||
| + | Species are increasingly seen to be better modelled as individuals. | ||
| + | |||
| + | ===Advantages=== | ||
| + | * Taxa are modeled as individuals. | ||
| + | * Taxonomic rank terms are ordered through their ''part_of'' relationships. | ||
| + | * Multiple part_of parents introduce fewer problems in inference than multiple is_a parents. This is important as more cases of reticulate phylogeny, caused by, for example, cross species hybridization or horizontal gene transfer, are discovered. | ||
| + | |||
| + | ===Inference with an ontology of taxonomic ranks + instance graph of taxa=== | ||
| + | |||
| + | ===Individuals/Instances=== | ||
| + | Implementing this approach will require [[Tool support for Instances]]. | ||
| + | |||
| + | ==Taxonomic Ranks as Metaclasses== | ||
| + | |||
| + | An approach that allows taxa to be viewed as both classes and individuals is to view rank terms, such as 'genus' as metaclasses. A metaclass is a class of classes. This would mean that a particular genus, e.g., ''Danio'' is both a subclass of Cyprinidae and an instance of genus. Metaclasses are used when assigning particular properties to a class itself, rather than its members. The property 'extinct' might be such a class-level property, as opposed to 'dead' a property that inheres in individuals. Extinct is a property we want to inhere in certain taxa only when all their individual constituents are dead. However, there are classes, such as anatomical parts, or behavior patterns that would not be subject to the possiblity of extinction. | ||
| + | |||
| + | This approach is attractive because it allows taxa to be both classes, in line with traditional taxonomies and reasoning with description logic classifiers, and individuals, with the greater fidelity to current biological thinking. | ||
| + | |||
| + | ===Disadvantages=== | ||
| + | The main disadvantage with this approach is its complexity, both for human understanding and for reasoning. | ||
[[Category:Ontology]] | [[Category:Ontology]] | ||
[[Category:Taxonomy]] | [[Category:Taxonomy]] | ||
Revision as of 16:16, 31 August 2010
The Teleost Taxonomy Ontology (TTO) is based on the Catalog of Fishes and will be updated as CoF updates appear as well as from input from area experts. The reminder of this page will discuss how the TTO is currently implemented. See the Taxonomic Rank Ontology, an alternative implementation based on phylogenetic principles.
Contents
Organization
The current organization is derived from the ontology generated from the NCBI taxonomy. There is a hierarchy of taxon names, each of which is a class.
Cyprinidae --------------------> Family
^ has_rank
|
is_a |
|
Davario --------------------> Genus
^ has_rank
|
is_a |
|
Davario aequipinnatus ----------------> Species
has_rank
Resources
Here are some useful resources related to the taxonomy ontology (TTO)
- Change list (relative to November 2007 CoF update) for TTO. This list is current for recent changes - older changes will be added.
- Suggested ideas for taxonomy browser and other tools
- Summary of a discussion of TTO issues at the second data jamboree (September 2008)
This is a list of resources related to the development of the TTO.
- Chris Mungall's NCBI Taxonomy in OBO format - this file is huge!
- Catalog of Fishes database schema documentation. - possibly somewhat out of date
- OBO Flat File Format Specification
- OBO Identifier Lifecycle
Taxonomy ontology structure
There is now a page devoted to discussion of Taxonomic Ranks.
The NCBI taxonomy ontology has some special terms and relations which are probably relevant to the fish taxonomy ontology. One of these is the "has_rank" typedef, which is used as a property value on the taxon terms:
[Typedef] id: has_rank name: has_rank def: "A metadata relation between a class and its taxonomic rank (eg species, family)" [] comment: This is an abstract class for use with the NCBI taxonomy to name the depth of the node within the tree. The link between the node term and the rank is only visible if you are using an obo 1.3 aware browser/editor; otherwise this can be ignored is_metadata_tag: true
Some special terms are included which represent taxonomic ranks. They descend from the term "taxonomic_rank":
[Term] id: NCBITaxon:taxonomic_rank name: taxonomic_rank def: "A level of depth within a species taxonomic tree" [] comment: This is an abstract class for use with the NCBI taxonomy to name the depth of the node within the tree. The link between the node term and the rank is only visible if you are using an obo 1.3 aware browser/editor; otherwise this can be ignored [Term] id: NCBITaxon:superkingdom name: superkingdom is_a: NCBITaxon:taxonomic_rank [Term] id: NCBITaxon:genus name: genus is_a: NCBITaxon:taxonomic_rank [Term] id: NCBITaxon:species name: species is_a: NCBITaxon:taxonomic_rank etc....
Here are three actual taxon terms. The term for the species descends via is_a from the term for the genus, and likewise for the genus and family. The rank is specified in the property_value. Other names are placed in the synonym fields:
[Term] id: NCBITaxon:7996 name: Ictaluridae is_a: NCBITaxon:7995 synonym: "North American freshwater catfishes" EXACT common_name [] synonym: "bullhead catfishes" EXACT genbank_common_name [] property_value: has_rank NCBITaxon:family xref: GC_ID:1 [Term] id: NCBITaxon:7997 name: Ictalurus is_a: NCBITaxon:7996 property_value: has_rank NCBITaxon:genus xref: GC_ID:1 [Term] id: NCBITaxon:7998 name: Ictalurus punctatus is_a: NCBITaxon:7997 synonym: "channel catfish" EXACT genbank_common_name [] property_value: has_rank NCBITaxon:species xref: GC_ID:1
Other uses for the synonym field can be seen in the OBO file, such as for obsolete names. We may want to also provide a definition from the original species description.
Taxonomy ontology content from Catalog of Fishes
Which parts of the CoF schema should be included in the taxonomy ontology? Also, is there information not in the Catalog of Fishes that should be included in the taxonomy ontology?
There is now a rough draft Taxonomy ontology directly from (somewhat dated) dump files from the Catalog of Fishes.
A draft version with synonyms is here. The actual top of the tree is still Craniata, other species
appearing at the root represent problems - usually related to synonyms that we need to resolve. Also note that I have tagged all
synonyms as "RELATED", the weakest category of synonymy. Given the complexities of taxonomic synonymy, this seems to be the appropriate
place to start. With the most recent version, most synonyms also include database references (dbxrefs) back to the original genus and species id's from the CoF tables.
Here is a preliminary taxonomy ontology, using data from a somewhat dated CoF dump. The taxa represented here are mentioned in
Fink, S.V., and W. L. Fink. 1981. Interrelationships of the ostariophysan fishes (Teleostei). Zoological Journal of the Linnean
Society 72: 297–353.
Craniata is included as a root for the entire taxonomy, even though it does not correspond to any taxonomic level defined in the ontology.
Note also that the term ids are not generated by OBO edit, we are basing them on catalog numbers from CoF.
Craniata: 1 Our root
Class: 10 + the class number from CoF (are these subject to splits?)
Order: 1000 + 10 * the order number from CoF - these numbers are sometimes formatted XX.X (to indicate splits?)
Family: 10000 + 10 * the family number from CoF - these numbers are sometimes formatted XX.X (to indicate splits?)
Genus: 100000 + the CAS_GEN value from CoF
Species: 1000000 + the CAS_SPC value from CoF
Unnamed entities: 10000000+ (e.g., sp., spp., etc.)
We should probably discuss whether this id scheme is really what we want to do - it means we can't add taxa from OBOEdit, though that isn't necessarily a problem.
format-version: 1.2 date: 02:10:2007 15:27 saved-by: peter auto-generated-by: OBO-Edit 1.101 default-namespace: teleost-taxonomy [Term] id: TTO: 1029899 name: Prochilodus vimboides is_a: TTO:101208 ! Prochilodus property_value: has_rank TTO:species [Term] id: TTO:1 name: Craniata [Term] id: TTO:10000001 name: Eigenmannia sp. is_a: TTO:104678 ! Eigenmannia property_value: has_rank TTO:Unknown_species [Term] id: TTO:10000002 name: Sternopygus sp. is_a: TTO:102060 ! Sternopygus property_value: has_rank TTO:Unknown_species [Term] id: TTO:1003114 name: Notemigonus crysoleucus is_a: TTO:100967 ! Notemigonus property_value: has_rank TTO:species [Term] id: TTO:1003239 name: Opsariichthys uncirostris is_a: TTO:103159 ! Opsariichthys property_value: has_rank TTO:species [Term] id: TTO:1004143 name: Chalceus macrolepidotus is_a: TTO:100825 ! Chalceus property_value: has_rank TTO:species [Term] id: TTO:1004205 name: Hepsetus odoe is_a: TTO:109162 ! Hepsetus property_value: has_rank TTO:species [Term] id: TTO:1004213 name: Xenocharax spilurus is_a: TTO:103438 ! Xenocharax property_value: has_rank TTO:species [Term] id: TTO:1004343 name: Rhoadsia altipinna is_a: TTO:105589 ! Rhoadsia property_value: has_rank TTO:species [Term] id: TTO:100558 name: Chanos is_a: TTO:10720 ! Chanidae property_value: has_rank TTO:genus [Term] id: TTO:1006103 name: Rhabdolichops troscheli is_a: TTO:107098 ! Rhabdolichops property_value: has_rank TTO:species [Term] id: TTO:100825 name: Chalceus is_a: TTO:10910 ! Characidae property_value: has_rank TTO:genus [Term] id: TTO:100967 name: Notemigonus is_a: TTO:10760 ! Cyprinidae property_value: has_rank TTO:genus [Term] id: TTO:101208 name: Prochilodus is_a: TTO:10855 ! Prochilodontidae property_value: has_rank TTO:genus [Term] id: TTO:101842 name: Brycon is_a: TTO:10910 ! Characidae property_value: has_rank TTO:genus [Term] id: TTO:102060 name: Sternopygus is_a: TTO:11231 ! Sternopygidae property_value: has_rank TTO:genus [Term] id: TTO:1027661 name: Chanos chanos is_a: TTO:100558 ! Chanos property_value: has_rank TTO:species [Term] id: TTO:1028504 name: Bryconamericus brevirostris is_a: TTO:105385 ! Bryconamericus property_value: has_rank TTO:species [Term] id: TTO:102915 name: Diplomystes is_a: TTO:10920 ! Diplomystidae property_value: has_rank TTO:genus [Term] id: TTO:1029536 name: Brycon dentex is_a: TTO:101842 ! Brycon property_value: has_rank TTO:species [Term] id: TTO:1030110 name: Platyurosternarchus macrostomus synonym: "Sternarchus macrostoma" EXACT [] is_a: TTO:110391 ! Platyurosternarchus property_value: has_rank TTO:species [Term] id: TTO:103159 name: Opsariichthys is_a: TTO:10760 ! Cyprinidae property_value: has_rank TTO:genus [Term] id: TTO:103339 name: Auchenoglanis is_a: TTO:10940 ! Bagridae property_value: has_rank TTO:genus [Term] id: TTO:103438 name: Xenocharax is_a: TTO:10820 ! Citharinidae property_value: has_rank TTO:genus [Term] id: TTO:1046341 name: Diplomystes chilensis synonym: "Diplomystes papillosus" EXACT [] is_a: TTO:102915 ! Diplomystes property_value: has_rank TTO:species [Term] id: TTO:104678 name: Eigenmannia is_a: TTO:11231 ! Sternopygidae property_value: has_rank TTO:genus [Term] id: TTO:105055 name: Zacco is_a: TTO:10760 ! Cyprinidae property_value: has_rank TTO:genus [Term] id: TTO:1051779 name: Zacco temminckii is_a: TTO:105055 ! Zacco property_value: has_rank TTO:species [Term] id: TTO:105268 name: Sternarchorhamphus is_a: TTO:11232 ! Apteronotidae property_value: has_rank TTO:genus [Term] id: TTO:105385 name: Bryconamericus is_a: TTO:10910 ! Characidae property_value: has_rank TTO:genus [Term] id: TTO:105589 name: Rhoadsia is_a: TTO:10910 ! Characidae property_value: has_rank TTO:genus [Term] id: TTO:1056213 name: Auchenoglanis monkei synonym: "Parauchenoglanis guttatus" EXACT [] is_a: TTO:103339 ! Auchenoglanis property_value: has_rank TTO:species [Term] id: TTO:1062171 name: Sternopygus macrurus is_a: TTO:102060 ! Sternopygus property_value: has_rank TTO:species [Term] id: TTO:107098 name: Rhabdolichops is_a: TTO:11231 ! Sternopygidae property_value: has_rank TTO:genus [Term] id: TTO:10720 name: Chanidae is_a: TTO:1350 ! Gonorynchiformes property_value: has_rank TTO:family [Term] id: TTO:10760 name: Cyprinidae is_a: TTO:1360 ! Cypriniformes property_value: has_rank TTO:family [Term] id: TTO:10820 name: Citharinidae is_a: TTO:1370 ! Characiformes property_value: has_rank TTO:family [Term] id: TTO:10827 name: Hepsetidae is_a: TTO:1370 ! Characiformes property_value: has_rank TTO:family [Term] id: TTO:10855 name: Prochilodontidae is_a: TTO:1370 ! Characiformes property_value: has_rank TTO:family [Term] id: TTO:10910 name: Characidae is_a: TTO:1370 ! Characiformes property_value: has_rank TTO:family [Term] id: TTO:109162 name: Hepsetus is_a: TTO:10827 ! Hepsetidae property_value: has_rank TTO:genus [Term] id: TTO:10920 name: Diplomystidae is_a: TTO:1380 ! Siluriformes property_value: has_rank TTO:family [Term] id: TTO:10940 name: Bagridae is_a: TTO:1380 ! Siluriformes property_value: has_rank TTO:family [Term] id: TTO:110391 name: Platyurosternarchus is_a: TTO:11232 ! Apteronotidae property_value: has_rank TTO:genus [Term] id: TTO:11231 name: Sternopygidae is_a: TTO:1390 ! Gymnotiformes property_value: has_rank TTO:family [Term] id: TTO:11232 name: Apteronotidae is_a: TTO:1390 ! Gymnotiformes property_value: has_rank TTO:family [Term] id: TTO:1350 name: Gonorynchiformes is_a: TTO:18 ! Actinopterygii property_value: has_rank TTO:order [Term] id: TTO:1360 name: Cypriniformes is_a: TTO:18 ! Actinopterygii property_value: has_rank TTO:order [Term] id: TTO:1370 name: Characiformes is_a: TTO:18 ! Actinopterygii property_value: has_rank TTO:order [Term] id: TTO:1380 name: Siluriformes is_a: TTO:18 ! Actinopterygii property_value: has_rank TTO:order [Term] id: TTO:1390 name: Gymnotiformes is_a: TTO:18 ! Actinopterygii property_value: has_rank TTO:order [Term] id: TTO:18 name: Actinopterygii is_a: TTO:1 ! Craniata property_value: has_rank TTO:class [Term] id: TTO:class name: class xref: NCBITaxon:class is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:family name: family xref: NCBITaxon:family is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:genus name: genus xref: NCBITaxon:genus is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:order name: order xref: NCBITaxon:order is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:species name: species xref: NCBITaxon:species is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:taxonomic_rank name: taxonomic_rank xref: NCBITaxon:taxonomic_rank [Term] id: TTO:Unknown_species name: Unknown species is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:Unknown_species_group name: Unknown species group is_a: TTO:taxonomic_rank ! taxonomic_rank [Term] id: TTO:Unnamed_species name: Unnamed species is_a: TTO:taxonomic_rank ! taxonomic_rank
Note: This page describes some design alternatives for taxonomic ontologies that refer to or include terms for ranks. If you are looking documentation for the Vocabulary of Ranks developed by Phenoscape, you will find it here.
A taxonomic rank ontology allows particular taxa to be treated as historical individuals instead of as universal types.
Previous attempts to represent taxonomy using ontology usually include taxonomic groups as classes in the ontology. Individual organisms are seen as instances of those universal types. There could be an ontology term Mammal, such that Primates and Rodents are more particular types of Mammals (is_a descendants). Taxonomies are even often used as examples to help explain ontology inheritance to new users. This scheme fails to represent reality in several ways and, as this page will demonstrate, is even misleading.
The "traditional" view
Inference with traditional taxonomy
One of the attractions of a traditional taxonomy for ontology builders that it can support the most common sort of reasoning for Description Logic ontologies. Although neither the TTO nor the NCBI taxonomy ontology attach character values to their classes, in principle such characters could be extracted from keys or from the taxonomic literature. With the addition of such properties, individuals can be classified by starting at the root of the taxonomy and traversing down the tree until either a terminal taxon (species) is reached or a node is reached where no descendant has the combination of characters displayed by the individual. For biologists, this operation is identical to use of an identification key. Such an operation is generally only defined for is_a links.
Problems with the "traditional" view
- The only criterion for inclusion within a taxonomic group is physical descent from a member of that group (we are assuming a taxonomy consistent with phylogeny). There are no universal properties one could ascribe to members of "Mammalia", besides things the members happen to share at this moment. "Hair" is often used an example of a Mammalian property; but of course its commonality is the result of its presence in the common ancestor of all mammals, and many mammals lack hair almost entirely. A mammal species that had no hair at all would nevertheless still be a mammal. Such reversals are common in phylogenies and can occur for both morphological properties (e.g., loss of legs in snakes) and molecular characters (e.g., inferred origin of pseudogenes).
- Descendant terms do not inherit (ontologically) all of the characteristics of their parents. For example, Rodentia has a different age and geographic range from Mammalia.
- Properties of a taxonomic group change through time: its geographic range, genetic and morphological diversity, etc. It can go extinct. These properties inhere in the group, not any one individual.
- Treating taxonomic groups as "classes" or "types" reinforces essentialist ideas of life forms, which evolutionary biology has completely overturned. Essentialist thought is a hindrance to understanding of the evolutionary process.
- Most contemporary treatments of the species concept (refs) treat them, and in many cases higher taxa, as individuals, possessing a history.
Instead of treating taxa as types in an ontology, we can treat individual taxa as instances of the type Taxon, or more specifically as instances of some taxonomic rank (see figure above).
Taxonomic Rank Ontology + Instance Graph of Taxonomy
Species are increasingly seen to be better modelled as individuals.
Advantages
- Taxa are modeled as individuals.
- Taxonomic rank terms are ordered through their part_of relationships.
- Multiple part_of parents introduce fewer problems in inference than multiple is_a parents. This is important as more cases of reticulate phylogeny, caused by, for example, cross species hybridization or horizontal gene transfer, are discovered.
Inference with an ontology of taxonomic ranks + instance graph of taxa
Individuals/Instances
Implementing this approach will require Tool support for Instances.
Taxonomic Ranks as Metaclasses
An approach that allows taxa to be viewed as both classes and individuals is to view rank terms, such as 'genus' as metaclasses. A metaclass is a class of classes. This would mean that a particular genus, e.g., Danio is both a subclass of Cyprinidae and an instance of genus. Metaclasses are used when assigning particular properties to a class itself, rather than its members. The property 'extinct' might be such a class-level property, as opposed to 'dead' a property that inheres in individuals. Extinct is a property we want to inhere in certain taxa only when all their individual constituents are dead. However, there are classes, such as anatomical parts, or behavior patterns that would not be subject to the possiblity of extinction.
This approach is attractive because it allows taxa to be both classes, in line with traditional taxonomies and reasoning with description logic classifiers, and individuals, with the greater fidelity to current biological thinking.
Disadvantages
The main disadvantage with this approach is its complexity, both for human understanding and for reasoning.